Also, even though some studies have uncovered several similarities, there are others that have detected notable differences between archaeal and eukaryotic TBP. This sequence was originally called Box A, which is now known to be the sequence that interacts with the homologue of the archaeal TATA-binding protein (TBP). In archaea species, the promoter contains an 8 bp AT-rich sequence located ~24 bp upstream of the transcription start site. 
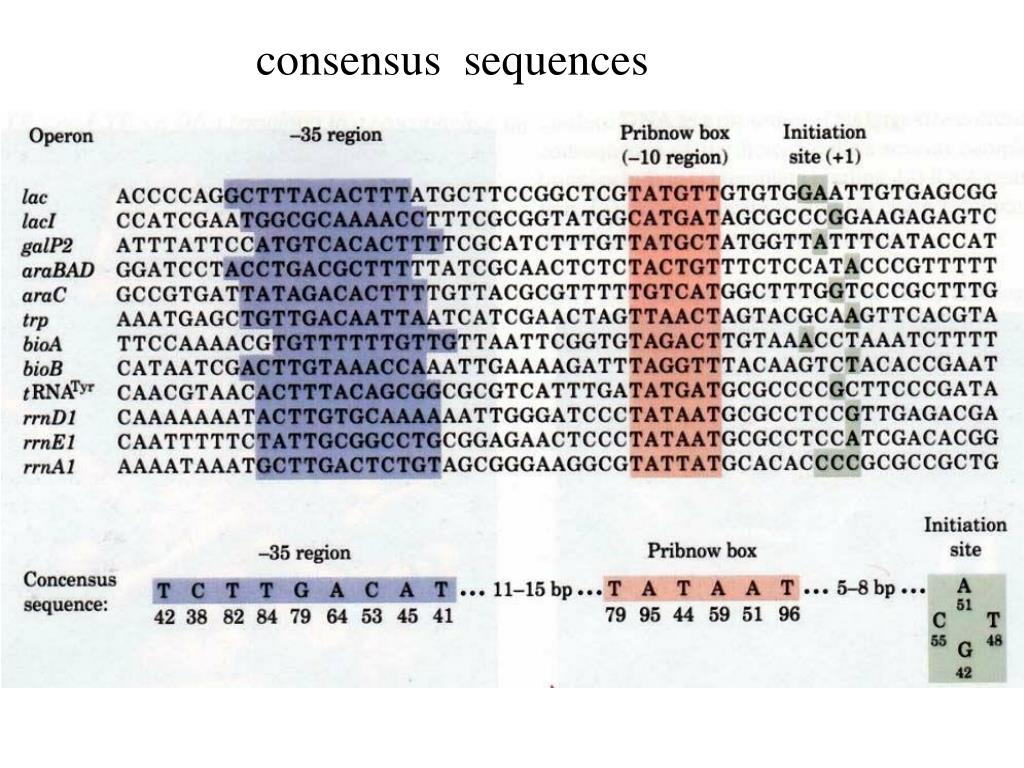
Most research on the TATA box has been conducted on yeast, human, and Drosophila genomes, however, similar elements have been found in archaea and ancient eukaryotes. The TATA box was found in protein coding genes transcribed by RNA polymerase II. They first discovered the TATA sequence while analyzing 5' DNA promoter sequences in Drosophila, mammalian, and viral genes. The TATA box was the first eukaryotic core promoter motif to be identified in 1978 by American biochemist David Hogness while he and his graduate student, Michael Goldberg were on sabbatical at the University of Basel in Switzerland. The TATA-binding protein (TBP) could also be targeted by viruses as a means of viral transcription. Some diseases associated with mutations in the TATA box include gastric cancer, spinocerebellar ataxia, Huntington's disease, blindness, β-thalassemia, immunosuppression, Gilbert's syndrome, and HIV-1. These phenotypic changes can then turn into a disease phenotype. Without proper regulation of transcription, eukaryotic organisms would not be able to properly respond to their environment.īased on the sequence and mechanism of TATA box initiation, mutations such as insertions, deletions, and point mutations to this consensus sequence can result in phenotypic changes. Gene transcription by RNA polymerase II depends on the regulation of the core promoter by long-range regulatory elements such as enhancers and silencers. The TATA box is the binding site of the TATA-binding protein (TBP) and other transcription factors in some eukaryotic genes. Transcription is initiated at the TATA box in TATA-containing genes. The TATA box was first identified in 1978 as a component of eukaryotic promoters. The boxing in of sequences sheds light on the origin of the term "box". When consensus nucleotides and alternative ones were compared, homologous regions were "boxed" by the researchers. In the 1980s, while investigating nucleotide sequences in mouse genome loci, the Hogness box sequence was found and "boxed in" at the -31 position. How the term "box" originated is unclear. It was termed the "TATA box" as it contains a consensus sequence characterized by repeating T and A base pairs. The TATA box is considered a non-coding DNA sequence (also known as a cis-regulatory element). The bacterial homolog of the TATA box is called the Pribnow box which has a shorter consensus sequence. In molecular biology, the TATA box (also called the Goldberg–Hogness box) is a sequence of DNA found in the core promoter region of genes in archaea and eukaryotes. The TATA box consensus sequence is TATAWAW, where W is either A or T.
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
